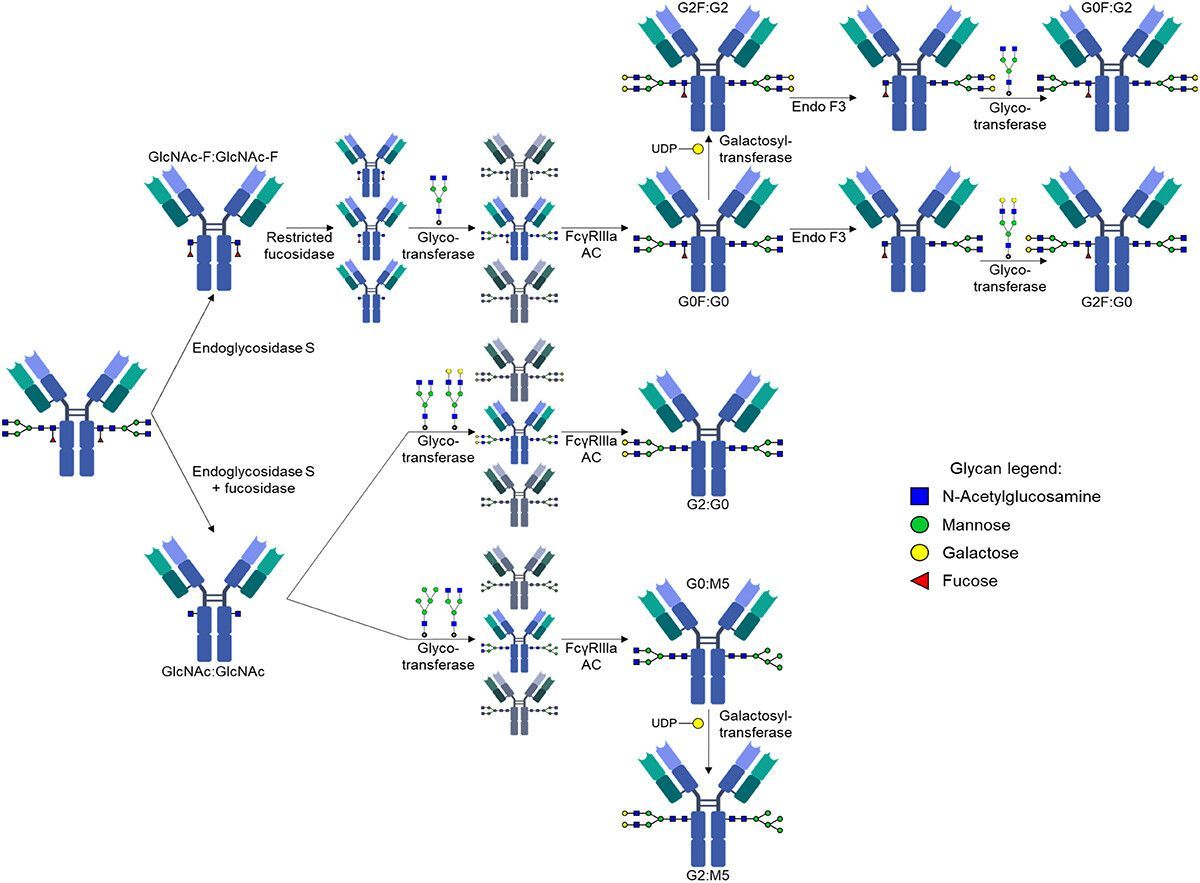
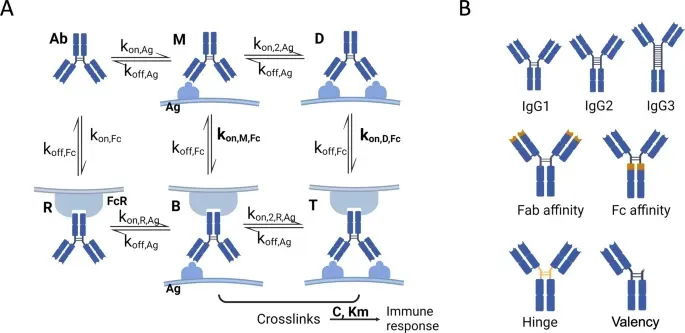
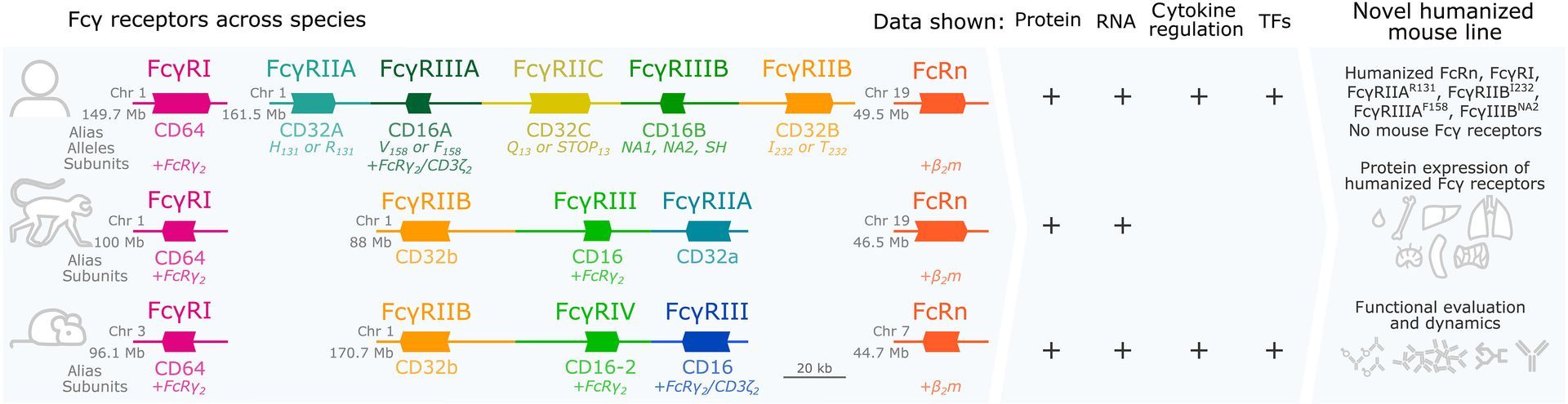
From early discovery through clinical development and post-licensure, SeromYx’s Systems Serology platform helps vaccine developers understand the functional immune mechanisms that drive protection.
We support natural infection studies, controlled human infection models, field efficacy trials, and post-licensure analyses by identifying functional correlates of protection and immune signatures linked to efficacy. Our data informs critical decisions around antigen and adjuvant selection, dose optimization, manufacturing process changes, and comparison of vaccine candidates against licensed products.
Pathogens Studied Using Systems Serology
Our Systems Serology platform has been applied across a wide range of infectious diseases to support vaccine development, immunogenicity assessment, and immune correlate discovery. The pathogens below represent viruses, bacteria, parasites, and fungi that have been studied using SeromYx’s comprehensive Fc-mediated antibody profiling and functional immune analysis.
DISCOVERY
Systems Serology identifies antibody features linked to disease outcomes.
Profiling hospitalized COVID-19 patients revealed distinct antibody responses: survivors showed a spike (S)-focused, functional profile, while non-survivors had nucleocapsid (N)-biased, less effective antibodies. Just five key antibody features predicted clinical trajectory, supporting early risk stratification and biomarker development. These findings highlight the importance of spike-specific functional antibodies for vaccine targeting.
- Differentiates survivors and non-survivors using key antibody signatures
- Links enhanced S-specific phagocytosis and complement activity to protection
- Informs vaccine antigen selection and immune monitoring strategies
LEAD SELECTION
Prioritize candidates by protective antibody functions, not just quantity.
Systems Serology enables comparison of vaccine regimens based on their ability to elicit key protective antibody functions. Screening HIV vaccine candidates revealed wide variation in ADCP levels despite similar antibody titers. Changes in adjuvants and dosing shifted functional profiles, helping developers select candidates that induce higher-quality immune responses. This accelerates confident, data-driven down-selection for clinical advancement.
- Differentiates vaccine leads by antibody function, not just titer
- Reveals impact of formulation and regimen on immune quality
- Supports efficient selection of candidates with greater clinical potential
ASSAY DEVELOPMENT, VALIDATION, QUALIFICATION
Robust immune profiling beyond neutralization for clinical trials.
SeromYx developed and qualified a Systems Serology assay under WHO and GCLP standards to measure antibody-dependent functions in SARS-CoV-2 vaccine trials. The assay delivers precise, specific, and sensitive measurement of Fc effector activities (ADCP, ADCD, ADNKA), supporting secondary endpoints in Phase 3 studies and regulatory filings. This validated platform is adaptable for diverse vaccine programs requiring advanced immune monitoring.
- Ensures reproducible, regulatory-grade immune data across cohorts
- Measures extra-neutralizing Fc functions linked to protection
- Scalable qualification framework for broad vaccine applications
CLINICAL EFFICACY TESTING
Uncovering antibody mechanisms linked to protection beyond neutralization.
A head-to-head study of mRNA-1273 and BNT162b2 vaccines showed both induced strong humoral responses, but mRNA-1273 elicited higher IgA titers, enhanced Fcγ receptor binding, and greater functional activity (ADNP, ADNKA). Systems Serology mapped distinct Fc effector functions, offering mechanistic insights into their differing real-world efficacy and supporting identification of immune correlates of protection.
- Differentiates vaccine-induced functional antibody profiles linked to protection
- Detects spike-specific Fc activities beyond neutralization
- Provides mechanistic insight for clinical trial correlates and disease attenuation
MANUFACTURING SCALE-UP
Integrating antibody avidity with Fc effector profiling to enhance vaccine assessment.
By combining Systems Serology with surface plasmon resonance (SPR)–based avidity measurements, this study uncovered distinct immune profiles differentiating two vaccine manufacturing processes. While one process drove higher effector function with lower avidity, the other generated stronger avidity with reduced effector activity. Multivariate modeling highlighted avidity, NK cell IFNγ responses, and IgG1 as key features distinguishing the processes, providing mechanistic insight into how manufacturing influences vaccine-induced immunity.
- Links antibody avidity with functional immune readouts
- Differentiates immune profiles across vaccine manufacturing processes
- Provides mechanistic insight to guide vaccine design and optimization
Frequently Asked Questions (FAQ)
Can you analyze clinical trial samples?
Yes. Our platform supports preclinical through clinical-stage vaccine programs, including natural infection studies, controlled human infection models, and field efficacy trials.
Can you identify immune signatures associated with protection?
Yes. Using systems-level analytics, we identify multivariate immune features that associate with protection, durability, or reduced disease severity.
When is the best time to engage SeromYx in a program?
We provide value across all phases, but especially during:
- Candidate down-selection
- Phase I/II immunogenicity studies
- Correlate of protection investigations
What are the sample requirements for vaccine studies?
Sample volume requirements vary depending on the assay panel selected. In general, we require approximately 25–150 µL per assay when performed in isolation against a single antigen.
When multiple assays are combined within an integrated panel, overall sample requirements may decrease due to optimized assay design.
If sample volume is limited, particularly for clinical trial cohorts, we can develop a tailored strategy to maximize data generation while conserving material. We encourage teams to contact us for a study-specific estimate as it may fall outside the range provided above.